
Amino Acid Residue-driven Nanoparticle Targeting of Protein Cavities Beyond Size Complementarity
Cavities at protein-protein interaction interfaces are often considered "undruggable" because their shallow or large geometries hinder the stable binding by small molecules. A study published in Journal of the American Chemical Society and led by Prof. LI Yang from the Shenzhen Institutes of Advanced Technology (SIAT) of the Chinese Academy of Sciences elucidated the molecular mechanisms governing nanoparticles (NPs) recognition and selective targeting of protein surface cavities.
Using SARS-CoV-2 S trimer (S trimer) as a model system, researchers compared CeO2NPs and gold nanoparticles (AuNPs) to evaluate the role of surface chemistry in protein cavity targeting, which have comparable sizes but different surface chemistries. They found that both NPs interacted with the S trimer and exhibited antiviral effects, but differed in their preferred binding sites and molecular mechanisms.
In detail, CeO2NPs bound to the central cavity enriched in aspartic acid (Asp) residues where coordination with Asp carboxyl groups enabled stable binding and interfered with ACE2 receptor recognition. In contrast, AuNPs predominantly bound to arginine (Arg)-rich lateral cavities near the S1/S2 cleavage region through electrostatic interactions and hydrogen bonding, thereby interfering with host protease-mediated activation of the S trimer.
Furthermore, researchers showed that NP targeting of protein cavities could not be explained by geometric accessibility alone. Instead, the selectivity depended on how NP surface chemistry matched the local amino acid environment within a cavity. Precise cavity recognition therefore required both geometric compatibility and surface chemical interactions.
The findings of this study improve the understanding of how NPs selectively bind to protein surface cavities, and provide guidance for the rational design of NPs to modulate protein function.
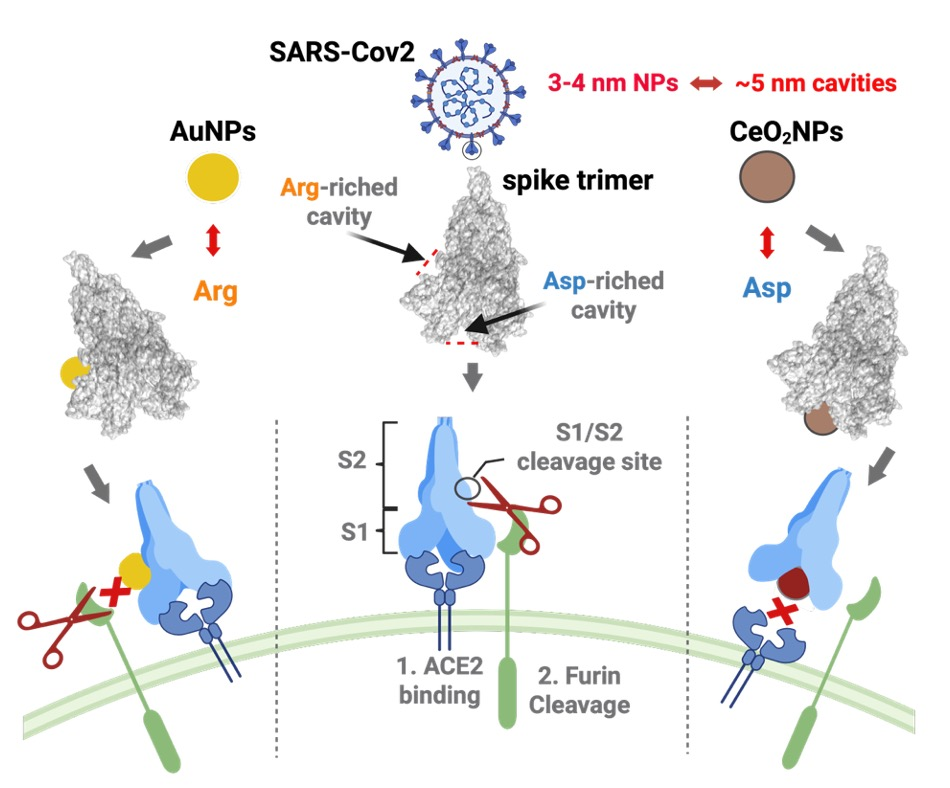
Amino acid residue-driven binding of size-matched NPs to protein cavities of the SARS-CoV-2 S trimer.(Image by SIAT)
File Download: